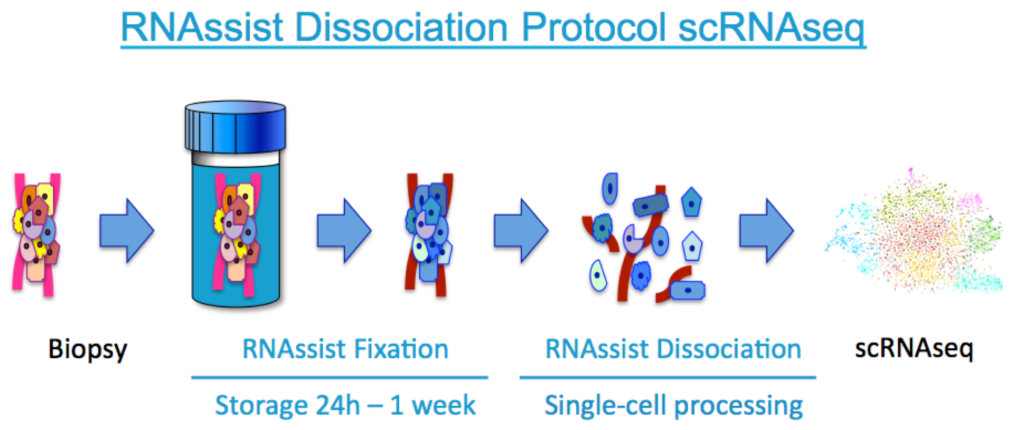
vivoPHIX™ is a universal platform technology for preparing higher quality single cells for scRNA-seq (patent pending). The technology enables the immediate stabilisation of the transcriptome at the point of sampling, tissue fixation, cell dissociation, pathogen inactivation and qualitatively improved scRNA-seq results. Advantageously, either entire pieces of tissue or single-cells can be stored and transported prior to further use and scRNA-seq analysis.
The combination of a novel mechanical methodology with a non-enzymatic, non-aqueous tissue dissociation buffer fully protects the integrity of nucleic acids and proteins from the initial point of sample fixation throughout the entire dissociation procedure providing high quality single-cells for scRNA-seq.
Leading academic laboratories, including the Sanger and Pasteur Institutes have demonstrated that a broad range of tissues types including insect (mosquito) and mammal (mouse and human) can be efficiently processed into single cells.
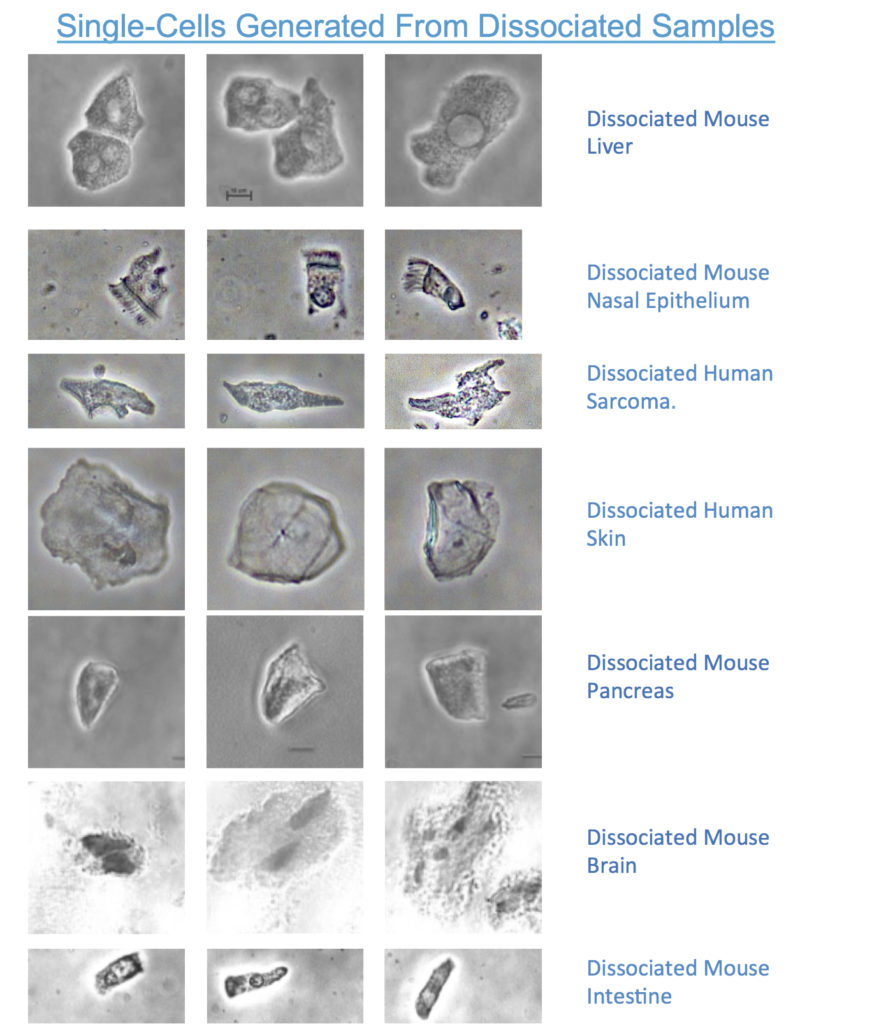
Tissue dissociation has been demonstrated for intestine, nasal epithelium, kidney, liver, pancreas, skin and a PDX sample. Unprecedentedly, such dissociated cells retain their original morphology such as 3D shape, brush borders and dendrites, providing a means to select cells based on in vivo shape, as well as providing metadata for scRNA-seq and other downstream molecular analytical techniques.
The combination of a novel type of efficient mechanical dissociation, protective buffer and simplified cellular capture reduces cell clumps, doublets and cell debris. Because the cellular transcriptome is immediately stabilised by vivoPHIX at the point of sampling (e.g. fresh biopsy) and retains the protective environment until single-cells are captured and transferred into the aqueous scRNA-seq front-end buffer.
We believe that a great deal of the original in situ cellular morphology is retained during tissue dissociation leading to cells with unique morphologies potentially providing unique insights distinguishing healthy from diseased tissues.
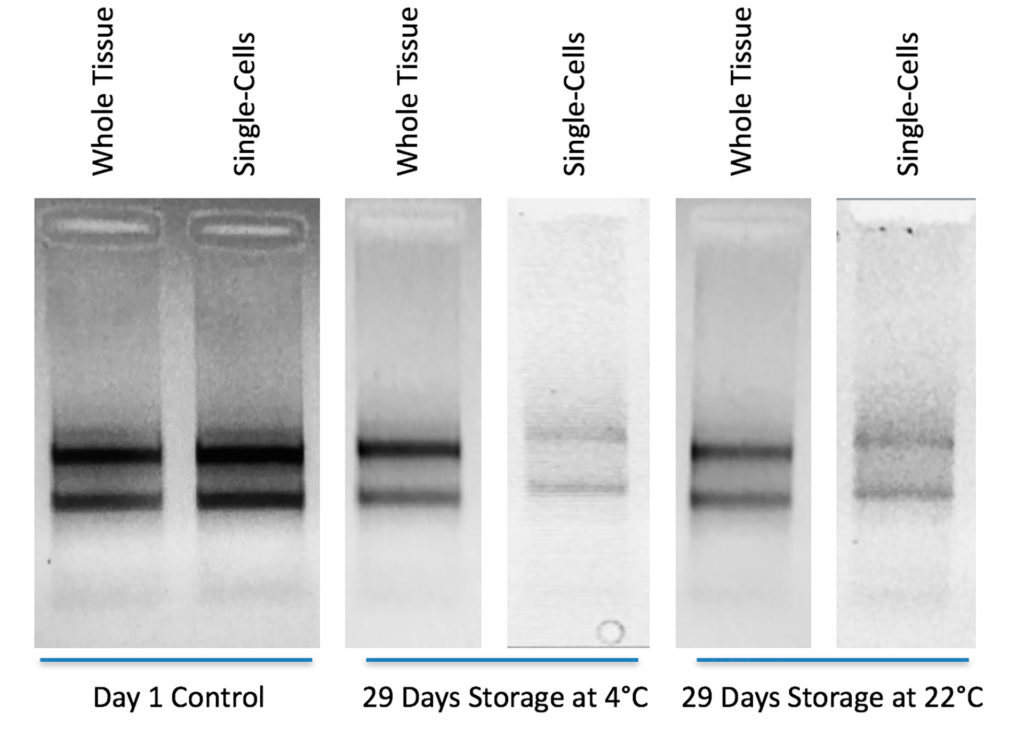
Total RNA was extracted (RNeasy, Qiagen) from bulk liver tissue immediately after a 2 hour fixation period. A second portion of the bulk tissue was then stored at either 4°C or 20°C for 29 days before RNA extraction. A third portion of the liver tissue was dissociated with the vivoPHIX dissociation protocol (patent-pending) and the single-cells stored at either 4°C or 20°C for 29 days before bulk RNA extraction on the single-cells. Gel electrophoresis of the RNA demonstrates that RNA degradation of the single-cells and bulk liver was minimal. This demonstrates that it is technically possible to store single-cells until such a time as is appropriate to run a scRNA-seq experiment, opening up the possibility of running multiple samples together rather than on a one-by-one as available basis.